KBQA知识图谱问答
1.学习图数据库Neo4j
1.1引言
知识图谱数据包含实体、属性、关系等,存储一般采用图数据库(Graph Databases),而Neo4j是其中最为常见的图数据库。官网下载对应系统的Neo4j。
cypher:是Neo4J的声明式图形查询语言,允许用户不必编写图形结构的遍历代码,就可以对图形数据进行高效的查询。
1.2创建节点
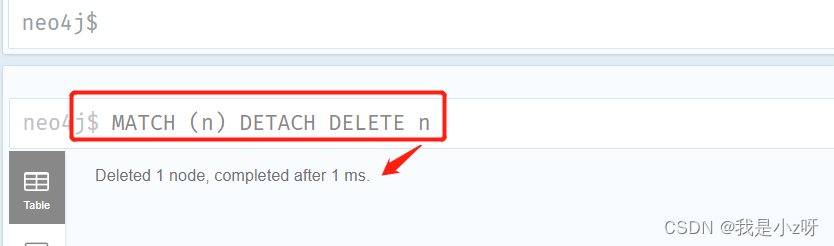
1.打开neo4j中一个库的browser,删除库中的数据
MATCH (n) DETACH DELETE n
MATCH是匹配操作,而小括号()代表一个节点node(可理解为括号类似一个圆形),括号里面的n为标识符。
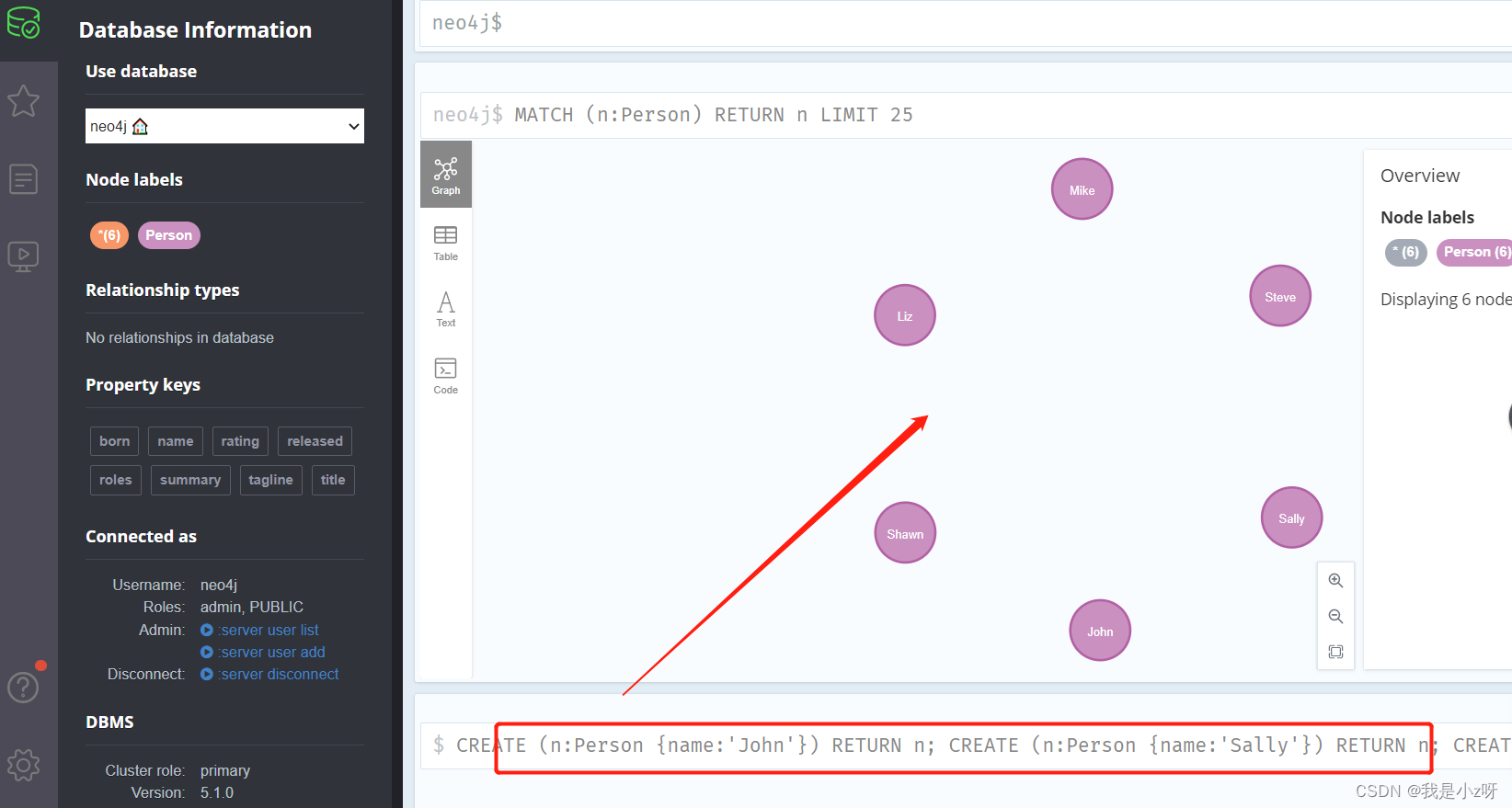
2.创建6个人物节点
CREATE (n:Person {
name:'John'}) RETURN n;
CREATE (n:Person {
name:'Sally'}) RETURN n;
CREATE (n:Person {
name:'Steve'}) RETURN n;
CREATE (n:Person {
name:'Mike'}) RETURN n;
CREATE (n:Person {
name:'Liz'}) RETURN n;
CREATE (n:Person {
name:'Shawn'}) RETURN n
CREATE是创建操作,Person是标签,代表节点的类型;花括号{}代表节点的属性,属性类似Python的字典;这条语句的含义就是创建一个标签为Person的节点,该节点具有一个name属性,属性值是John。
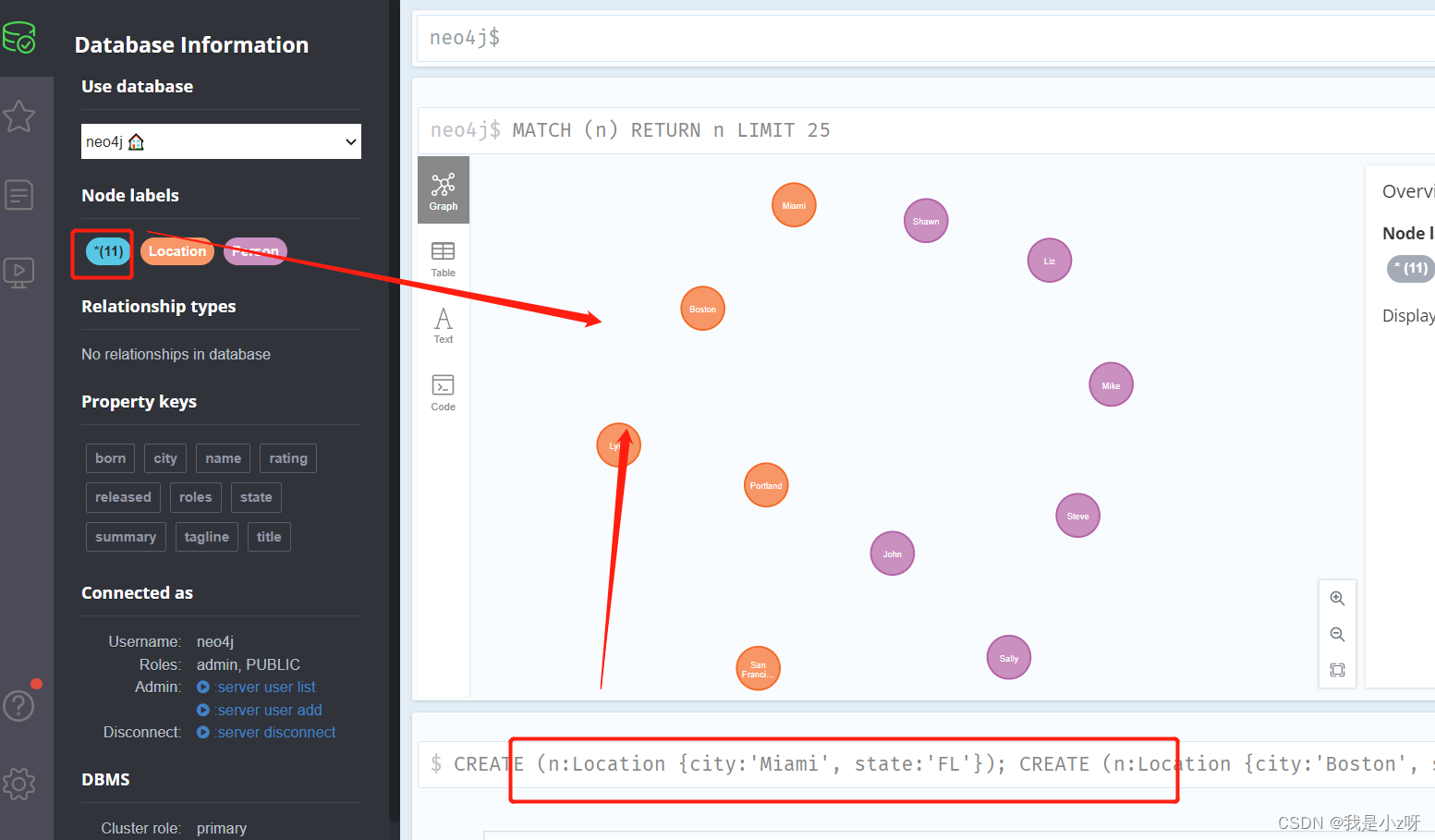
3.创建5个地区节点
CREATE (n:Location {
city:'Miami', state:'FL'});
CREATE (n:Location {
city:'Boston', state:'MA'});
CREATE (n:Location {
city:'Lynn', state:'MA'});
CREATE (n:Location {
city:'Portland', state:'ME'});
CREATE (n:Location {
city:'San Francisco', state:'CA'})
节点类型为Location,属性包括city和state。
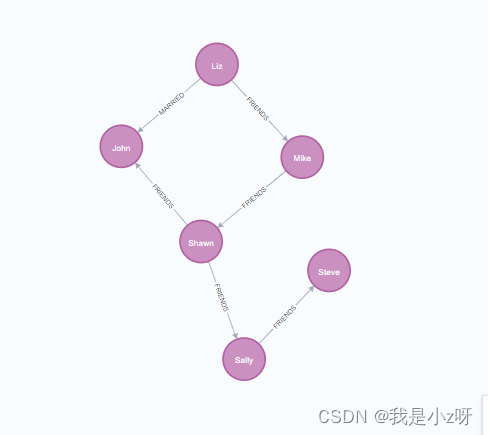
1.3创建关系
1.同种类型创建关系
MATCH (a:Person{
name:'Liz'}),(b:Person{
name:'Mike'}) MERGE (a)-[:FRIENDS]->(b);
MATCH (a:Person {
name:'Shawn'}), (b:Person {
name:'John'}) MERGE (a)-[:FRIENDS {
since:2012}]->(b);
MATCH (a:Person {
name:'Mike'}), (b:Person {
name:'Shawn'}) MERGE (a)-[:FRIENDS {
since:2006}]->(b);
MATCH (a:Person {
name:'Sally'}), (b:Person {
name:'Steve'}) MERGE (a)-[:FRIENDS {
since:2006}]->(b);
MATCH (a:Person {
name:'Liz'}), (b:Person {
name:'John'}) MERGE (a)-[:MARRIED {
since:1998}]->(b);
MATCH (a:Person {
name:'Shawn'}), (b:Person {
name:'Sally'}) MERGE (a)-[:FRIENDS since:2001}]->(b)
方括号[]即为关系,FRIENDS为关系的类型;注意这里的箭头–>是有方向的,表示是从a到b的关系。 这样,Liz和Mike之间建立了FRIENDS关系;{}在关系[]中是给关系增加属性。
2.不同类型创建关系
MATCH (a:Person {
name:'John'}), (b:Location {
city:'Boston'}) MERGE (a)-[:BORN_IN {
year:1978}]->(b);
MATCH (a:Person {
name:'Liz'}), (b:Location {
city:'Boston'}) MERGE (a)-[:BORN_IN {
year:1981}]->(b);
MATCH (a:Person {
name:'Mike'}), (b:Location {
city:'San Francisco'}) MERGE (a)-[:BORN_IN {
year:1960}]->(b);
MATCH (a:Person {
name:'Shawn'}), (b:Location {
city:'Miami'}) MERGE (a)-[:BORN_IN {
year:1960}]->(b);
MATCH (a:Person {
name:'Steve'}), (b:Location {
city:'Lynn'}) MERGE (a)-[:BORN_IN {
year:1970}]->(b)
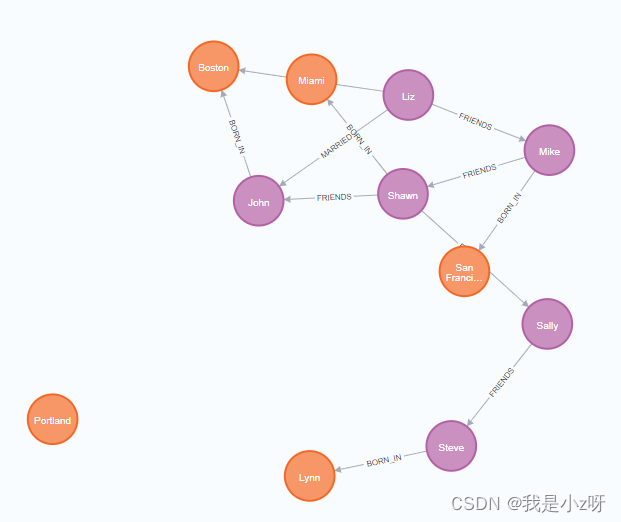
3.创建节点时就建立关系
CREATE (a:Person {
name:'Todd'})-[r:FRIENDS]->(b:Person {
name:'Carlos'})
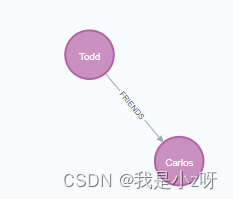
1.4查询
1.查询下所有在Boston出生的人物
MATCH (a:Person)-[:BORN_IN]->(b:Location {
city:'Boston'}) RETURN a,b
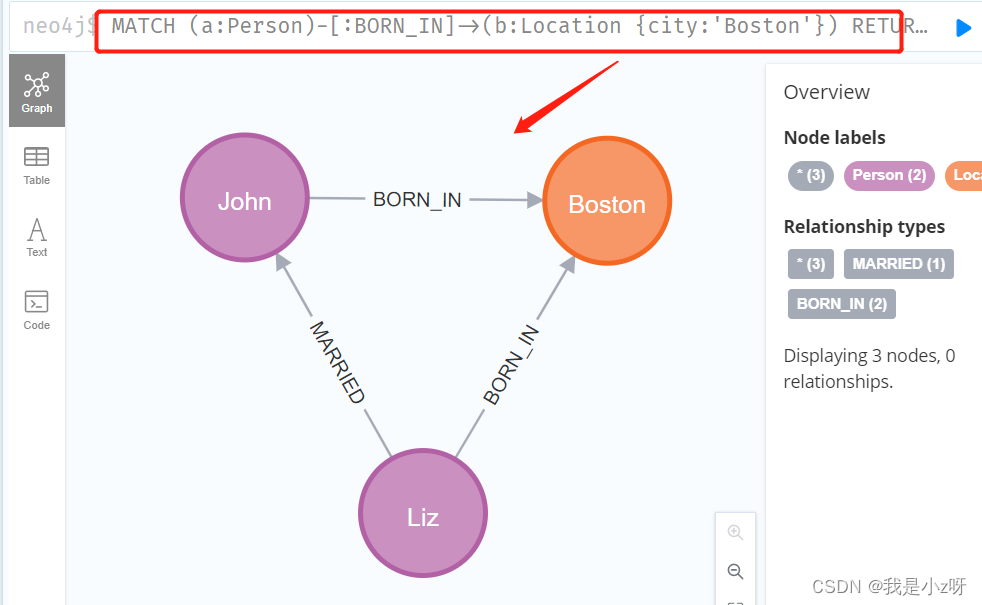
2. 查询所有有关系的节点
MATCH (a)--() RETURN a
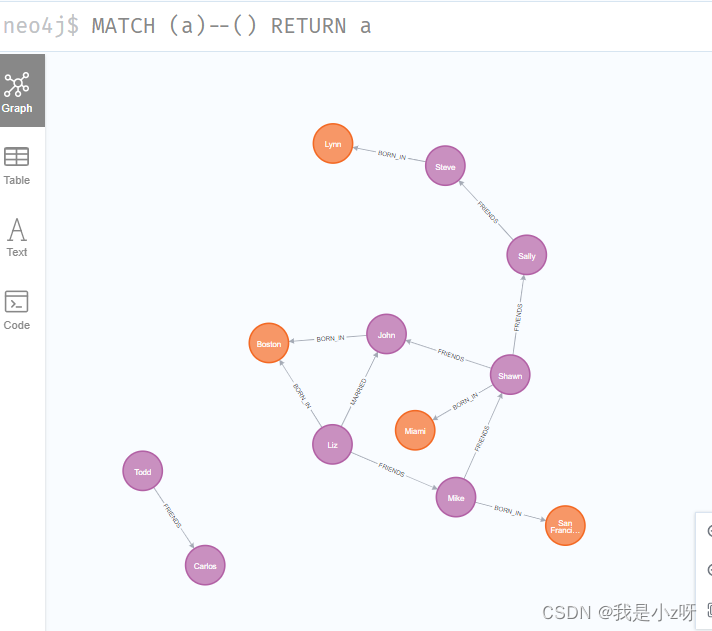
3.查询所有对外有关系的节点(这里有箭头,表明了关系方向是向外)
MATCH (a)-->() RETURN a
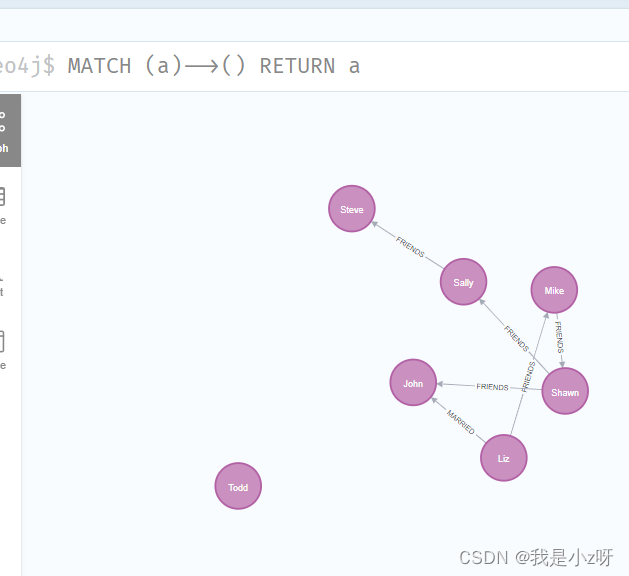
4.查询所有对外有关系的节点,以及关系类型
MATCH (a)-[r]->() RETURN a.name, type(r)
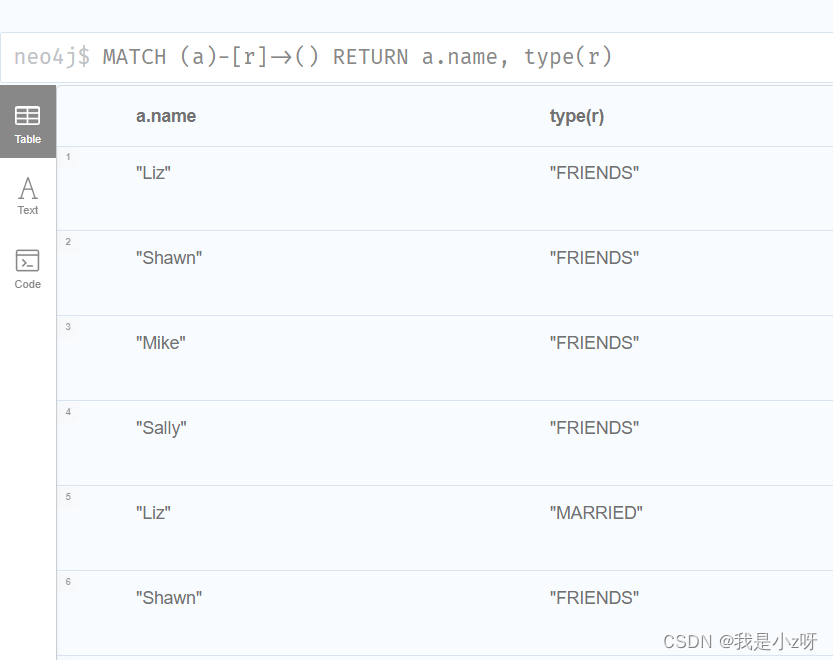
5.查询所有有结婚关系的节点
MATCH (n)-[:MARRIED]-() RETURN n
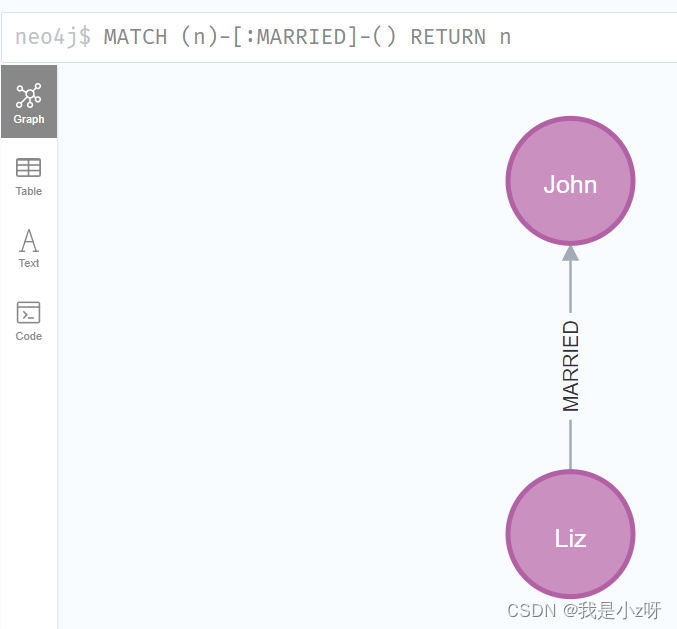
6.查找某人的朋友的朋友
MATCH (a:Person {
name:'Mike'})-[r1:FRIENDS]-()-[r2:FRIENDS]-(friend_of_a_friend) RETURN friend_of_a_friend.name AS fofName
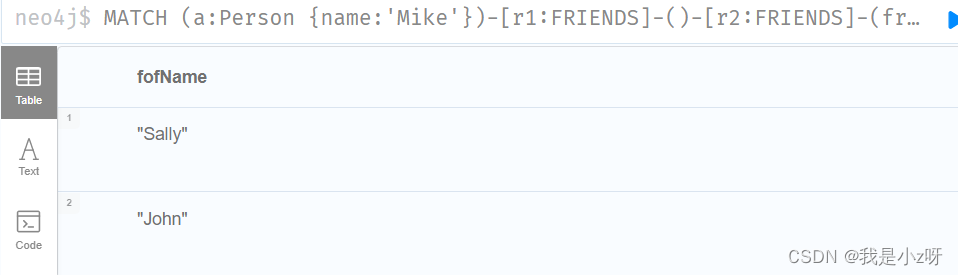
1.5修改和删除
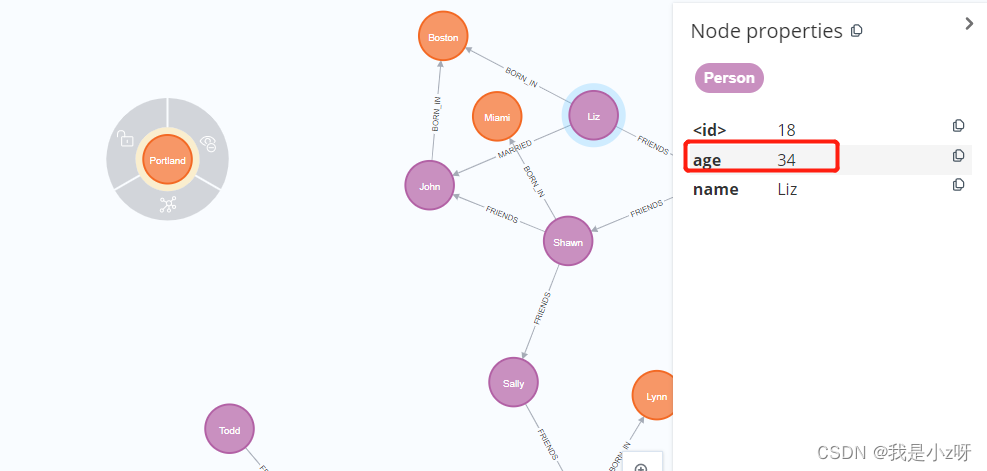
1.增加/修改节点属性
MATCH (a:Person {
name:'Liz'}) SET a.age=34;
MATCH (a:Person {
name:'Shawn'}) SET a.age=32;
MATCH (a:Person {
name:'John'}) SET a.age=44;
MATCH (a:Person {
name:'Mike'}) SET a.age=25
SET表示修改操作
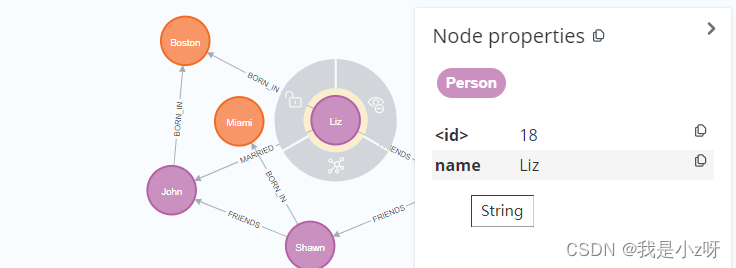
2.删除节点的属性
MATCH (a:Person {
name:'Liz'}) REMOVE a.age
REMOVE表示删除节点属性操作
3.删除节点
MATCH (a:Location {
city:'Portland'}) DELETE a
DELETE是删除节点操作
4.删除有关系的节点
MATCH (a:Person {
name:'Todd'})-[rel]-(b:Person) DELETE a,b,rel
2.医药知识图谱问答
这里我用了刘焕勇老师github项目中的医药知识图谱自动问答系统进行学习。
2.1 搭建项目需要的python虚拟环境。
conda create -n kbqa python
conda activate kbqa
pip install lxml
pip install pymongdb
pip install py2neo
pip install pyahocorasick -i HTTPS://mirrors.aliyun.com/pypi/simple/
安装mongdb,参考博文。
安装neo4j,参考博文。neo4j和java的版本要对应,不然运行neo4j console时会出现错误Run with ‘–verbose’ for a more detailed error message,这里可以参考博文和博文。这里我搞了好久,其实就两个问题,给环境变量里面path里面加入java的bin目录和neo4j的bin目录;这样就可以cmd中查看这两个的version,然后就是我运行neo4j console时出现了Please use Oracle® Java™ 17, OpenJDK™ 17 to run Neo4j,这句话的意思就是java要用17版本的,而不是别的版本的,这里注意看自己需要哪个版本,我因为只看博文换了好多版本,后来才明白看这个报错已经明确说了要17版。
还有就是一直报错== main ERROR Cannot access RandomAccessFile java.io.FileNotFoundException: D:\Softwares\Neo4j\neo4j-community-5.2.0\logs\neo4j.log (拒绝访问。) ==这个拒绝访问就是权限不够,我把文件都放到了c盘的program files文件夹下了。
StatusLogger Unable to create file D:\Program Files\neo4j-community-4.4.0\logs\debug.log,说明该路径没有权限,修改该文件夹权限为可读写。
2.2 运行data_spider.py,爬取网页数据保存到mongodb。
data_spider.py
#!/usr/bin/env python3
# coding: utf-8
import urllib.request
import urllib.parse
from lxml import etree
import pymongo
import re
class CrimeSpider:
def __init__(self):
self.conn = pymongo.MongoClient()
self.db = self.conn['medical']#库
self.col = self.db['data']#表
'''根据url,请求html'''
def get_html(self, url):
headers = {
'User-Agent': 'Mozilla/5.0 (Windows NT 10.0; WOW64) AppleWebKit/537.36 (KHTML, like Gecko) '
'Chrome/51.0.2704.63 Safari/537.36'}
req = urllib.request.Request(url=url, headers=headers)
res = urllib.request.urlopen(req)
html = res.read().decode('gbk')
return html
'''url解析'''
def url_parser(self, content):
selector = etree.HTML(content)
urls = ['http://www.anliguan.com' + i for i in selector.xpath('//h2[@class="item-title"]/a/@href')]
return urls
'''测试'''
def spider_main(self):
for page in range(1, 100):
try:
basic_url = 'http://jib.xywy.com/il_sii/gaishu/%s.htm'%page
cause_url = 'http://jib.xywy.com/il_sii/cause/%s.htm'%page
prevent_url = 'http://jib.xywy.com/il_sii/prevent/%s.htm'%page
symptom_url = 'http://jib.xywy.com/il_sii/symptom/%s.htm'%page
inspect_url = 'http://jib.xywy.com/il_sii/inspect/%s.htm'%page
treat_url = 'http://jib.xywy.com/il_sii/treat/%s.htm'%page
food_url = 'http://jib.xywy.com/il_sii/food/%s.htm'%page
drug_url = 'http://jib.xywy.com/il_sii/drug/%s.htm'%page
data = {
}
data['url'] = basic_url
data['basic_info'] = self.basicinfo_spider(basic_url)
data['cause_info'] = self.common_spider(cause_url)
data['prevent_info'] = self.common_spider(prevent_url)
data['symptom_info'] = self.symptom_spider(symptom_url)
data['inspect_info'] = self.inspect_spider(inspect_url)
data['treat_info'] = self.treat_spider(treat_url)
data['food_info'] = self.food_spider(food_url)
data['drug_info'] = self.drug_spider(drug_url)
print(page, basic_url)
self.col.insert_one(data)
except Exception as e:
print(e, page)
return
'''基本信息解析'''
def basicinfo_spider(self, url):
html = self.get_html(url)
selector = etree.HTML(html)
title = selector.xpath('//title/text()')[0]
category = selector.xpath('//div[@class="wrap mt10 nav-bar"]/a/text()')
desc = selector.xpath('//div[@class="jib-articl-con jib-lh-articl"]/p/text()')
ps = selector.xpath('//div[@class="mt20 articl-know"]/p')
infobox = []
for p in ps:
info = p.xpath('string(.)').replace('\r','').replace('\n','').replace('\xa0', '').replace(' ', '').replace('\t','')
infobox.append(info)
basic_data = {
}
basic_data['category'] = category
basic_data['name'] = title.split('的简介')[0]
basic_data['desc'] = desc
basic_data['attributes'] = infobox
return basic_data
'''treat_infobox治疗解析'''
def treat_spider(self, url):
html = self.get_html(url)
selector = etree.HTML(html)
ps = selector.xpath('//div[starts-with(@class,"mt20 articl-know")]/p')
infobox = []
for p in ps:
info = p.xpath('string(.)').replace('\r','').replace('\n','').replace('\xa0', '').replace(' ', '').replace('\t','')
infobox.append(info)
return infobox
'''treat_infobox治疗解析'''
def drug_spider(self, url):
html = self.get_html(url)
selector = etree.HTML(html)
drugs = [i.replace('\n','').replace('\t', '').replace(' ','') for i in selector.xpath('//div[@class="fl drug-pic-rec mr30"]/p/a/text()')]
return drugs
'''food治疗解析'''
def food_spider(self, url):
html = self.get_html(url)
selector = etree.HTML(html)
divs = selector.xpath('//div[@class="diet-img clearfix mt20"]')
try:
food_data = {
}
food_data['good'] = divs[0].xpath('./div/p/text()')
food_data['bad'] = divs[1].xpath('./div/p/text()')
food_data['recommand'] = divs[2].xpath('./div/p/text()')
except:
return {
}
return food_data
'''症状信息解析'''
def symptom_spider(self, url):
html = self.get_html(url)
selector = etree.HTML(html)
symptoms = selector.xpath('//a[@class="gre" ]/text()')
ps = selector.xpath('//p')
detail = []
for p in ps:
info = p.xpath('string(.)').replace('\r','').replace('\n','').replace('\xa0', '').replace(' ', '').replace('\t','')
detail.append(info)
symptoms_data = {
}
symptoms_data['symptoms'] = symptoms
symptoms_data['symptoms_detail'] = detail
return symptoms, detail
'''检查信息解析'''
def inspect_spider(self, url):
html = self.get_html(url)
selector = etree.HTML(html)
inspects = selector.xpath('//li[@class="check-item"]/a/@href')
return inspects
'''通用解析模块'''
def common_spider(self, url):
html = self.get_html(url)
selector = etree.HTML(html)
ps = selector.xpath('//p')
infobox = []
for p in ps:
info = p.xpath('string(.)').replace('\r', '').replace('\n', '').replace('\xa0', '').replace(' ','').replace('\t', '')
if info:
infobox.append(info)
return '\n'.join(infobox)
'''检查项抓取模块'''
def inspect_crawl(self):
for page in range(1, 36):
try:
url = 'http://jck.xywy.com/jc_%s.html'%page#这里是要爬取的网站链接
html = self.get_html(url)
data = {
}
data['url']= url
data['html'] = html
self.db['jc'].insert_one(data)
print(url)
except Exception as e:
print(e)
handler = CrimeSpider()
#抓取网页数据并保存到本地的mongodb数据库。库名叫medical:self.db = self.conn['medical']
handler.spider_main()#self.db['data'].insert_one('url','basic_info','cause_info','prevent_info'
#'symptom_info','inspect_info','treat_info','food_info','drug_info')中插入数据,
handler.inspect_crawl()#self.db['jc'].insert_one('url','html')中插入数据
报错:TypeError: ‘Collection’ object is not callable. If you meant to call the ‘insert’ method on a ‘Collection’ object it is failing because no such method exists。
原因:PyMongo 4.0.* 中 Collection.insert 方法被移除,使用 Collection.insert_one 或者 Collection.insert_many 替代。
解决:修改data_spider.py文件里的插入方法。
self.col.insert_one(data)
self.db['jc'].insert_one(data)
报错:localhost:27017: [WinError 10061] 由于目标计算机积极拒绝,无法连接。
原因:因为程序里链接了mongodb数据库,而我却没有开mongodb的后台。
解决:后台启动mongodb,在mongodb的bin目录下打开cdm
mongod.exe --dbpath D:\mongodb
在mongodb数据库里把数据导出成json文件。
2.3 运行build_medicalgraph.py文件,将json文件导入到neo4j数据库并抽取特征词。
#!/usr/bin/env python3
# coding: utf-8
import os
import json
from py2neo import Graph,Node
class MedicalGraph:
def __init__(self):
cur_dir = '/'.join(os.path.abspath(__file__).split('/')[:-1])
self.data_path = os.path.join(cur_dir, 'data/medical.json')
# self.g = Graph(
# host="127.0.0.1", # neo4j 搭载服务器的ip地址,ifconfig可获取到
# http_port=7474, # neo4j 服务器监听的端口号
# user="neo4j", # 数据库user name,如果没有更改过,应该是neo4j
# password="neo4j")
self.g = Graph('http://localhost:7474/', auth=('neo4j', '123qwe'))
'''读取文件'''
def read_nodes(self):
# 共7类节点(7+1)
drugs = [] # 药品
foods = [] # 食物
checks = [] # 检查
departments = [] #科室
producers = [] #药品大类
diseases = [] #疾病
symptoms = []#症状
disease_infos = []#疾病信息
# 构建节点实体关系(11)
rels_department = [] # 科室-科室关系
rels_noteat = [] # 疾病-忌吃食物关系
rels_doeat = [] # 疾病-宜吃食物关系
rels_recommandeat = [] # 疾病-推荐吃食物关系
rels_commonddrug = [] # 疾病-通用药品关系
rels_recommanddrug = [] # 疾病-热门药品关系
rels_check = [] # 疾病-检查关系
rels_drug_producer = [] # 厂商-药物关系
rels_symptom = [] #疾病症状关系
rels_acompany = [] # 疾病并发关系
rels_category = [] # 疾病与科室之间的关系
count = 0
for data in open(self.data_path,encoding='utf-8'):
disease_dict = {
}
count += 1
print(count)
data_json = json.loads(data)
disease = data_json['name']
disease_dict['name'] = disease
diseases.append(disease)
disease_dict['desc'] = ''
disease_dict['prevent'] = ''
disease_dict['cause'] = ''
disease_dict['easy_get'] = ''
disease_dict['cure_department'] = ''
disease_dict['cure_way'] = ''
disease_dict['cure_lasttime'] = ''
disease_dict['symptom'] = ''
disease_dict['cured_prob'] = ''
if 'symptom' in data_json:
symptoms += data_json['symptom']
for symptom in data_json['symptom']:
rels_symptom.append([disease, symptom])
if 'acompany' in data_json:
for acompany in data_json['acompany']:
rels_acompany.append([disease, acompany])
if 'desc' in data_json:
disease_dict['desc'] = data_json['desc']
if 'prevent' in data_json:
disease_dict['prevent'] = data_json['prevent']
if 'cause' in data_json:
disease_dict['cause'] = data_json['cause']
if 'get_prob' in data_json:
disease_dict['get_prob'] = data_json['get_prob']
if 'easy_get' in data_json:
disease_dict['easy_get'] = data_json['easy_get']
if 'cure_department' in data_json:
cure_department = data_json['cure_department']
if len(cure_department) == 1:
rels_category.append([disease, cure_department[0]])
if len(cure_department) == 2:
big = cure_department[0]
small = cure_department[1]
rels_department.append([small, big])
rels_category.append([disease, small])
disease_dict['cure_department'] = cure_department
departments += cure_department
if 'cure_way' in data_json:
disease_dict['cure_way'] = data_json['cure_way']
if 'cure_lasttime' in data_json:
disease_dict['cure_lasttime'] = data_json['cure_lasttime']
if 'cured_prob' in data_json:
disease_dict['cured_prob'] = data_json['cured_prob']
if 'common_drug' in data_json:
common_drug = data_json['common_drug']
for drug in common_drug:
rels_commonddrug.append([disease, drug])
drugs += common_drug
if 'recommand_drug' in data_json:
recommand_drug = data_json['recommand_drug']
drugs += recommand_drug
for drug in recommand_drug:
rels_recommanddrug.append([disease, drug])
if 'not_eat' in data_json:
not_eat = data_json['not_eat']
for _not in not_eat:
rels_noteat.append([disease, _not])
foods += not_eat
do_eat = data_json['do_eat']
for _do in do_eat:
rels_doeat.append([disease, _do])
foods += do_eat
recommand_eat = data_json['recommand_eat']
for _recommand in recommand_eat:
rels_recommandeat.append([disease, _recommand])
foods += recommand_eat
if 'check' in data_json:
check = data_json['check']
for _check in check:
rels_check.append([disease, _check])
checks += check
if 'drug_detail' in data_json:
drug_detail = data_json['drug_detail']
producer = [i.split('(')[0] for i in drug_detail]
rels_drug_producer += [[i.split('(')[0], i.split('(')[-1].replace(')', '')] for i in drug_detail]
producers += producer
disease_infos.append(disease_dict)
return set(drugs), set(foods), set(checks), set(departments), set(producers), set(symptoms), set(diseases), disease_infos,\
rels_check, rels_recommandeat, rels_noteat, rels_doeat, rels_department, rels_commonddrug, rels_drug_producer, rels_recommanddrug,\
rels_symptom, rels_acompany, rels_category
'''建立节点'''
def create_node(self, label, nodes):
count = 0
for node_name in nodes:
node = Node(label, name=node_name)
self.g.create(node)
count += 1
print(count, len(nodes))
return
'''创建知识图谱中心疾病的节点'''
def create_diseases_nodes(self, disease_infos):
count = 0
for disease_dict in disease_infos:
node = Node("Disease", name=disease_dict['name'], desc=disease_dict['desc'],
prevent=disease_dict['prevent'] ,cause=disease_dict['cause'],
easy_get=disease_dict['easy_get'],cure_lasttime=disease_dict['cure_lasttime'],
cure_department=disease_dict['cure_department']
,cure_way=disease_dict['cure_way'] , cured_prob=disease_dict['cured_prob'])
self.g.create(node)
count += 1
print(count)
return
'''创建知识图谱实体节点类型schema'''
def create_graphnodes(self):
Drugs, Foods, Checks, Departments, Producers, Symptoms, Diseases, disease_infos,rels_check, rels_recommandeat, rels_noteat, rels_doeat, rels_department, rels_commonddrug, rels_drug_producer, rels_recommanddrug,rels_symptom, rels_acompany, rels_category = self.read_nodes()
self.create_diseases_nodes(disease_infos)
self.create_node('Drug', Drugs)
print(len(Drugs))
self.create_node('Food', Foods)
print(len(Foods))
self.create_node('Check', Checks)
print(len(Checks))
self.create_node('Department', Departments)
print(len(Departments))
self.create_node('Producer', Producers)
print(len(Producers))
self.create_node('Symptom', Symptoms)
return
'''创建实体关系边'''
def create_graphrels(self):
Drugs, Foods, Checks, Departments, Producers, Symptoms, Diseases, disease_infos, rels_check, rels_recommandeat, rels_noteat, rels_doeat, rels_department, rels_commonddrug, rels_drug_producer, rels_recommanddrug,rels_symptom, rels_acompany, rels_category = self.read_nodes()
self.create_relationship('Disease', 'Food', rels_recommandeat, 'recommand_eat', '推荐食谱')
self.create_relationship('Disease', 'Food', rels_noteat, 'no_eat', '忌吃')
self.create_relationship('Disease', 'Food', rels_doeat, 'do_eat', '宜吃')
self.create_relationship('Department', 'Department', rels_department, 'belongs_to', '属于')
self.create_relationship('Disease', 'Drug', rels_commonddrug, 'common_drug', '常用药品')
self.create_relationship('Producer', 'Drug', rels_drug_producer, 'drugs_of', '生产药品')
self.create_relationship('Disease', 'Drug', rels_recommanddrug, 'recommand_drug', '好评药品')
self.create_relationship('Disease', 'Check', rels_check, 'need_check', '诊断检查')
self.create_relationship('Disease', 'Symptom', rels_symptom, 'has_symptom', '症状')
self.create_relationship('Disease', 'Disease', rels_acompany, 'acompany_with', '并发症')
self.create_relationship('Disease', 'Department', rels_category, 'belongs_to', '所属科室')
'''创建实体关联边'''
def create_relationship(self, start_node, end_node, edges, rel_type, rel_name):
count = 0
# 去重处理
set_edges = []
for edge in edges:
set_edges.append('###'.join(edge))
all = len(set(set_edges))
for edge in set(set_edges):
edge = edge.split('###')
p = edge[0]
q = edge[1]
query = "match(p:%s),(q:%s) where p.name='%s'and q.name='%s' create (p)-[rel:%s{name:'%s'}]->(q)" % (
start_node, end_node, p, q, rel_type, rel_name)
try:
self.g.run(query)
count += 1
print(rel_type, count, all)
except Exception as e:
print(e)
return
'''导出数据'''
def export_data(self):
Drugs, Foods, Checks, Departments, Producers, Symptoms, Diseases, disease_infos, rels_check, rels_recommandeat, rels_noteat, rels_doeat, rels_department, rels_commonddrug, rels_drug_producer, rels_recommanddrug, rels_symptom, rels_acompany, rels_category = self.read_nodes()
f_drug = open('drug.txt', 'w+')
f_food = open('food.txt', 'w+')
f_check = open('check.txt', 'w+')
f_department = open('department.txt', 'w+')
f_producer = open('producer.txt', 'w+')
f_symptom = open('symptoms.txt', 'w+')
f_disease = open('disease.txt', 'w+')
f_drug.write('\n'.join(list(Drugs)))
f_food.write('\n'.join(list(Foods)))
f_check.write('\n'.join(list(Checks)))
f_department.write('\n'.join(list(Departments)))
f_producer.write('\n'.join(list(Producers)))
f_symptom.write('\n'.join(list(Symptoms)))
f_disease.write('\n'.join(list(Diseases)))
f_drug.close()
f_food.close()
f_check.close()
f_department.close()
f_producer.close()
f_symptom.close()
f_disease.close()
return
if __name__ == '__main__':
handler = MedicalGraph()
print("step1:导入图谱节点中")
handler.create_graphnodes()
print("step2:导入图谱边中")
handler.create_graphrels()
#handler.export_data()#json>txt,抽取特证词
报错:ValueError: The following settings are not supported
原因:Py2neo默认为最新版本,版本太高。
解决:图链接数据库时用新的语法。
self.g = Graph('http://localhost:7474/', auth=("neo4j", "123qwe"))
报错:py2neo.errors.ConnectionUnavailable: (‘Cannot open connection to %r’, ConnectionProfile(‘http://localhost:7474’))
原因:没有启动neo4j
解决:
neo4j install-service
neo4j start
打开http://localhost:7474/
报错:‘gbk‘ codec can‘t decode byte 0xaf in position 81: illegal multibyte sequence
原因:文件的编码和解码的格式不一致
解决:
for data in open(self.data_path,encoding='utf-8'):
成功运行neo4j和build_medicalgraph.py文件后就可以在html中看到数据库图。

2.4 运行chatbot_graph.py文件,进行人机交互。
这里用到了3个文件:question_classifier.py:对可进行回答的问题进行分类;question_parser.py:对用户所提的问题进行解析并归属哪里问题;answer_search.py:对此问题在neo4j库中搜寻答案。
2.4.1 question_classifier.py 问题分类
#!/usr/bin/env python3
# coding: utf-8
import os
import ahocorasick
class QuestionClassifier:
def __init__(self):
cur_dir = '/'.join(os.path.abspath(__file__).split('/')[:-1])
# 特征词路径
self.disease_path = os.path.join(cur_dir, 'dict/disease.txt')
self.department_path = os.path.join(cur_dir, 'dict/department.txt')
self.check_path = os.path.join(cur_dir, 'dict/check.txt')
self.drug_path = os.path.join(cur_dir, 'dict/drug.txt')
self.food_path = os.path.join(cur_dir, 'dict/food.txt')
self.producer_path = os.path.join(cur_dir, 'dict/producer.txt')
self.symptom_path = os.path.join(cur_dir, 'dict/symptom.txt')
self.deny_path = os.path.join(cur_dir, 'dict/deny.txt')
# 加载特征词
self.disease_wds= [i.strip() for i in open(self.disease_path,encoding='utf-8') if i.strip()]
self.department_wds= [i.strip() for i in open(self.department_path,encoding='utf-8') if i.strip()]
self.check_wds= [i.strip() for i in open(self.check_path,encoding='utf-8') if i.strip()]
self.drug_wds= [i.strip() for i in open(self.drug_path,encoding='utf-8') if i.strip()]
self.food_wds= [i.strip() for i in open(self.food_path,encoding='utf-8') if i.strip()]
self.producer_wds= [i.strip() for i in open(self.producer_path,encoding='utf-8') if i.strip()]
self.symptom_wds= [i.strip() for i in open(self.symptom_path,encoding='utf-8') if i.strip()]
self.region_words = set(self.department_wds + self.disease_wds + self.check_wds + self.drug_wds + self.food_wds + self.producer_wds + self.symptom_wds)
self.deny_words = [i.strip() for i in open(self.deny_path,encoding='utf-8') if i.strip()]
# 构造领域actree
self.region_tree = self.build_actree(list(self.region_words))
# 构建词典
self.wdtype_dict = self.build_wdtype_dict()
# 问句疑问词
self.symptom_qwds = ['症状', '表征', '现象', '症候', '表现']
self.cause_qwds = ['原因','成因', '为什么', '怎么会', '怎样才', '咋样才', '怎样会', '如何会', '为啥', '为何', '如何才会', '怎么才会', '会导致', '会造成']
self.acompany_qwds = ['并发症', '并发', '一起发生', '一并发生', '一起出现', '一并出现', '一同发生', '一同出现', '伴随发生', '伴随', '共现']
self.food_qwds = ['饮食', '饮用', '吃', '食', '伙食', '膳食', '喝', '菜' ,'忌口', '补品', '保健品', '食谱', '菜谱', '食用', '食物','补品']
self.drug_qwds = ['药', '药品', '用药', '胶囊', '口服液', '炎片']
self.prevent_qwds = ['预防', '防范', '抵制', '抵御', '防止','躲避','逃避','避开','免得','逃开','避开','避掉','躲开','躲掉','绕开',
'怎样才能不', '怎么才能不', '咋样才能不','咋才能不', '如何才能不',
'怎样才不', '怎么才不', '咋样才不','咋才不', '如何才不',
'怎样才可以不', '怎么才可以不', '咋样才可以不', '咋才可以不', '如何可以不',
'怎样才可不', '怎么才可不', '咋样才可不', '咋才可不', '如何可不']
self.lasttime_qwds = ['周期', '多久', '多长时间', '多少时间', '几天', '几年', '多少天', '多少小时', '几个小时', '多少年']
self.cureway_qwds = ['怎么治疗', '如何医治', '怎么医治', '怎么治', '怎么医', '如何治', '医治方式', '疗法', '咋治', '怎么办', '咋办', '咋治']
self.cureprob_qwds = ['多大概率能治好', '多大几率能治好', '治好希望大么', '几率', '几成', '比例', '可能性', '能治', '可治', '可以治', '可以医']
self.easyget_qwds = ['易感人群', '容易感染', '易发人群', '什么人', '哪些人', '感染', '染上', '得上']
self.check_qwds = ['检查', '检查项目', '查出', '检查', '测出', '试出']
self.belong_qwds = ['属于什么科', '属于', '什么科', '科室']
self.cure_qwds = ['治疗什么', '治啥', '治疗啥', '医治啥', '治愈啥', '主治啥', '主治什么', '有什么用', '有何用', '用处', '用途',
'有什么好处', '有什么益处', '有何益处', '用来', '用来做啥', '用来作甚', '需要', '要']
print('model init finished ......')
return
'''分类主函数'''
def classify(self, question):
data = {
}
medical_dict = self.check_medical(question)
if not medical_dict:
return {
}
data['args'] = medical_dict
#收集问句当中所涉及到的实体类型
types = []
for type_ in medical_dict.values():
types += type_
question_type = 'others'
question_types = []
# 症状
if self.check_words(self.symptom_qwds, question) and ('disease' in types):
question_type = 'disease_symptom'
question_types.append(question_type)
if self.check_words(self.symptom_qwds, question) and ('symptom' in types):
question_type = 'symptom_disease'
question_types.append(question_type)
# 原因
if self.check_words(self.cause_qwds, question) and ('disease' in types):
question_type = 'disease_cause'
question_types.append(question_type)
# 并发症
if self.check_words(self.acompany_qwds, question) and ('disease' in types):
question_type = 'disease_acompany'
question_types.append(question_type)
# 推荐食品
if self.check_words(self.food_qwds, question) and 'disease' in types:
deny_status = self.check_words(self.deny_words, question)
if deny_status:
question_type = 'disease_not_food'
else:
question_type = 'disease_do_food'
question_types.append(question_type)
#已知食物找疾病
if self.check_words(self.food_qwds+self.cure_qwds, question) and 'food' in types:
deny_status = self.check_words(self.deny_words, question)
if deny_status:
question_type = 'food_not_disease'
else:
question_type = 'food_do_disease'
question_types.append(question_type)
# 推荐药品
if self.check_words(self.drug_qwds, question) and 'disease' in types:
question_type = 'disease_drug'
question_types.append(question_type)
# 药品治啥病
if self.check_words(self.cure_qwds, question) and 'drug' in types:
question_type = 'drug_disease'
question_types.append(question_type)
# 疾病接受检查项目
if self.check_words(self.check_qwds, question) and 'disease' in types:
question_type = 'disease_check'
question_types.append(question_type)
# 已知检查项目查相应疾病
if self.check_words(self.check_qwds+self.cure_qwds, question) and 'check' in types:
question_type = 'check_disease'
question_types.append(question_type)
# 症状防御
if self.check_words(self.prevent_qwds, question) and 'disease' in types:
question_type = 'disease_prevent'
question_types.append(question_type)
# 疾病医疗周期
if self.check_words(self.lasttime_qwds, question) and 'disease' in types:
question_type = 'disease_lasttime'
question_types.append(question_type)
# 疾病治疗方式
if self.check_words(self.cureway_qwds, question) and 'disease' in types:
question_type = 'disease_cureway'
question_types.append(question_type)
# 疾病治愈可能性
if self.check_words(self.cureprob_qwds, question) and 'disease' in types:
question_type = 'disease_cureprob'
question_types.append(question_type)
# 疾病易感染人群
if self.check_words(self.easyget_qwds, question) and 'disease' in types :
question_type = 'disease_easyget'
question_types.append(question_type)
# 若没有查到相关的外部查询信息,那么则将该疾病的描述信息返回
if question_types == [] and 'disease' in types:
question_types = ['disease_desc']
# 若没有查到相关的外部查询信息,那么则将该疾病的描述信息返回
if question_types == [] and 'symptom' in types:
question_types = ['symptom_disease']
# 将多个分类结果进行合并处理,组装成一个字典
data['question_types'] = question_types
return data
'''构造词对应的类型'''
def build_wdtype_dict(self):
wd_dict = dict()
for wd in self.region_words:
wd_dict[wd] = []
if wd in self.disease_wds:
wd_dict[wd].append('disease')
if wd in self.department_wds:
wd_dict[wd].append('department')
if wd in self.check_wds:
wd_dict[wd].append('check')
if wd in self.drug_wds:
wd_dict[wd].append('drug')
if wd in self.food_wds:
wd_dict[wd].append('food')
if wd in self.symptom_wds:
wd_dict[wd].append('symptom')
if wd in self.producer_wds:
wd_dict[wd].append('producer')
return wd_dict
'''构造actree,加速过滤'''
def build_actree(self, wordlist):
actree = ahocorasick.Automaton()
for index, word in enumerate(wordlist):
actree.add_word(word, (index, word))
actree.make_automaton()
return actree
'''问句过滤'''
def check_medical(self, question):
region_wds = []
for i in self.region_tree.iter(question):
wd = i[1][1]
region_wds.append(wd)
stop_wds = []
for wd1 in region_wds:
for wd2 in region_wds:
if wd1 in wd2 and wd1 != wd2:
stop_wds.append(wd1)
final_wds = [i for i in region_wds if i not in stop_wds]
final_dict = {
i:self.wdtype_dict.get(i) for i in final_wds}
return final_dict
'''基于特征词进行分类'''
def check_words(self, wds, sent):
for wd in wds:
if wd in sent:
return True
return False
if __name__ == '__main__':
handler = QuestionClassifier()
while 1:
question = input('input an question:')
data = handler.classify(question)
print(data)
2.4.2 question_parser.py 问题解析
#!/usr/bin/env python3
# coding: utf-8
class QuestionPaser:
'''构建实体节点'''
def build_entitydict(self, args):
entity_dict = {
}
for arg, types in args.items():
for type in types:
if type not in entity_dict:
entity_dict[type] = [arg]
else:
entity_dict[type].append(arg)
return entity_dict
'''解析主函数'''
def parser_main(self, res_classify):
args = res_classify['args']
entity_dict = self.build_entitydict(args)
question_types = res_classify['question_types']
sqls = []
for question_type in question_types:
sql_ = {
}
sql_['question_type'] = question_type
sql = []
if question_type == 'disease_symptom':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'symptom_disease':
sql = self.sql_transfer(question_type, entity_dict.get('symptom'))
elif question_type == 'disease_cause':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'disease_acompany':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'disease_not_food':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'disease_do_food':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'food_not_disease':
sql = self.sql_transfer(question_type, entity_dict.get('food'))
elif question_type == 'food_do_disease':
sql = self.sql_transfer(question_type, entity_dict.get('food'))
elif question_type == 'disease_drug':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'drug_disease':
sql = self.sql_transfer(question_type, entity_dict.get('drug'))
elif question_type == 'disease_check':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'check_disease':
sql = self.sql_transfer(question_type, entity_dict.get('check'))
elif question_type == 'disease_prevent':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'disease_lasttime':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'disease_cureway':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'disease_cureprob':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'disease_easyget':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
elif question_type == 'disease_desc':
sql = self.sql_transfer(question_type, entity_dict.get('disease'))
if sql:
sql_['sql'] = sql
sqls.append(sql_)
return sqls
'''针对不同的问题,分开进行处理'''
def sql_transfer(self, question_type, entities):
if not entities:
return []
# 查询语句
sql = []
# 查询疾病的原因
if question_type == 'disease_cause':
sql = ["MATCH (m:Disease) where m.name = '{0}' return m.name, m.cause".format(i) for i in entities]
# 查询疾病的防御措施
elif question_type == 'disease_prevent':
sql = ["MATCH (m:Disease) where m.name = '{0}' return m.name, m.prevent".format(i) for i in entities]
# 查询疾病的持续时间
elif question_type == 'disease_lasttime':
sql = ["MATCH (m:Disease) where m.name = '{0}' return m.name, m.cure_lasttime".format(i) for i in entities]
# 查询疾病的治愈概率
elif question_type == 'disease_cureprob':
sql = ["MATCH (m:Disease) where m.name = '{0}' return m.name, m.cured_prob".format(i) for i in entities]
# 查询疾病的治疗方式
elif question_type == 'disease_cureway':
sql = ["MATCH (m:Disease) where m.name = '{0}' return m.name, m.cure_way".format(i) for i in entities]
# 查询疾病的易发人群
elif question_type == 'disease_easyget':
sql = ["MATCH (m:Disease) where m.name = '{0}' return m.name, m.easy_get".format(i) for i in entities]
# 查询疾病的相关介绍
elif question_type == 'disease_desc':
sql = ["MATCH (m:Disease) where m.name = '{0}' return m.name, m.desc".format(i) for i in entities]
# 查询疾病有哪些症状
elif question_type == 'disease_symptom':
sql = ["MATCH (m:Disease)-[r:has_symptom]->(n:Symptom) where m.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
# 查询症状会导致哪些疾病
elif question_type == 'symptom_disease':
sql = ["MATCH (m:Disease)-[r:has_symptom]->(n:Symptom) where n.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
# 查询疾病的并发症
elif question_type == 'disease_acompany':
sql1 = ["MATCH (m:Disease)-[r:acompany_with]->(n:Disease) where m.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql2 = ["MATCH (m:Disease)-[r:acompany_with]->(n:Disease) where n.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql = sql1 + sql2
# 查询疾病的忌口
elif question_type == 'disease_not_food':
sql = ["MATCH (m:Disease)-[r:no_eat]->(n:Food) where m.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
# 查询疾病建议吃的东西
elif question_type == 'disease_do_food':
sql1 = ["MATCH (m:Disease)-[r:do_eat]->(n:Food) where m.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql2 = ["MATCH (m:Disease)-[r:recommand_eat]->(n:Food) where m.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql = sql1 + sql2
# 已知忌口查疾病
elif question_type == 'food_not_disease':
sql = ["MATCH (m:Disease)-[r:no_eat]->(n:Food) where n.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
# 已知推荐查疾病
elif question_type == 'food_do_disease':
sql1 = ["MATCH (m:Disease)-[r:do_eat]->(n:Food) where n.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql2 = ["MATCH (m:Disease)-[r:recommand_eat]->(n:Food) where n.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql = sql1 + sql2
# 查询疾病常用药品-药品别名记得扩充
elif question_type == 'disease_drug':
sql1 = ["MATCH (m:Disease)-[r:common_drug]->(n:Drug) where m.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql2 = ["MATCH (m:Disease)-[r:recommand_drug]->(n:Drug) where m.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql = sql1 + sql2
# 已知药品查询能够治疗的疾病
elif question_type == 'drug_disease':
sql1 = ["MATCH (m:Disease)-[r:common_drug]->(n:Drug) where n.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql2 = ["MATCH (m:Disease)-[r:recommand_drug]->(n:Drug) where n.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
sql = sql1 + sql2
# 查询疾病应该进行的检查
elif question_type == 'disease_check':
sql = ["MATCH (m:Disease)-[r:need_check]->(n:Check) where m.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
# 已知检查查询疾病
elif question_type == 'check_disease':
sql = ["MATCH (m:Disease)-[r:need_check]->(n:Check) where n.name = '{0}' return m.name, r.name, n.name".format(i) for i in entities]
return sql
if __name__ == '__main__':
handler = QuestionPaser()
2.4.3 answer_search.py 答案搜索
#!/usr/bin/env python3
# coding: utf-8
from py2neo import Graph
class AnswerSearcher:
def __init__(self):
# self.g = Graph(
# host="127.0.0.1",
# http_port=7474,
# user="neo4j",
# password="123qwe")
self.g = Graph('http://localhost:7474/', auth=("neo4j", "123qwe"))
self.num_limit = 20
'''执行cypher查询,并返回相应结果'''
def search_main(self, sqls):
final_answers = []
for sql_ in sqls:
question_type = sql_['question_type']
queries = sql_['sql']
answers = []
for query in queries:
ress = self.g.run(query).data()
answers += ress
final_answer = self.answer_prettify(question_type, answers)
if final_answer:
final_answers.append(final_answer)
return final_answers
'''根据对应的qustion_type,调用相应的回复模板'''
def answer_prettify(self, question_type, answers):
final_answer = []
if not answers:
return ''
if question_type == 'disease_symptom':
desc = [i['n.name'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}的症状包括:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'symptom_disease':
desc = [i['m.name'] for i in answers]
subject = answers[0]['n.name']
final_answer = '症状{0}可能染上的疾病有:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_cause':
desc = [i['m.cause'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}可能的成因有:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_prevent':
desc = [i['m.prevent'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}的预防措施包括:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_lasttime':
desc = [i['m.cure_lasttime'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}治疗可能持续的周期为:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_cureway':
desc = [';'.join(i['m.cure_way']) for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}可以尝试如下治疗:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_cureprob':
desc = [i['m.cured_prob'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}治愈的概率为(仅供参考):{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_easyget':
desc = [i['m.easy_get'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}的易感人群包括:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_desc':
desc = [i['m.desc'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0},熟悉一下:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_acompany':
desc1 = [i['n.name'] for i in answers]
desc2 = [i['m.name'] for i in answers]
subject = answers[0]['m.name']
desc = [i for i in desc1 + desc2 if i != subject]
final_answer = '{0}的症状包括:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_not_food':
desc = [i['n.name'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}忌食的食物包括有:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_do_food':
do_desc = [i['n.name'] for i in answers if i['r.name'] == '宜吃']
recommand_desc = [i['n.name'] for i in answers if i['r.name'] == '推荐食谱']
subject = answers[0]['m.name']
final_answer = '{0}宜食的食物包括有:{1}\n推荐食谱包括有:{2}'.format(subject, ';'.join(list(set(do_desc))[:self.num_limit]), ';'.join(list(set(recommand_desc))[:self.num_limit]))
elif question_type == 'food_not_disease':
desc = [i['m.name'] for i in answers]
subject = answers[0]['n.name']
final_answer = '患有{0}的人最好不要吃{1}'.format(';'.join(list(set(desc))[:self.num_limit]), subject)
elif question_type == 'food_do_disease':
desc = [i['m.name'] for i in answers]
subject = answers[0]['n.name']
final_answer = '患有{0}的人建议多试试{1}'.format(';'.join(list(set(desc))[:self.num_limit]), subject)
elif question_type == 'disease_drug':
desc = [i['n.name'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}通常的使用的药品包括:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'drug_disease':
desc = [i['m.name'] for i in answers]
subject = answers[0]['n.name']
final_answer = '{0}主治的疾病有{1},可以试试'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'disease_check':
desc = [i['n.name'] for i in answers]
subject = answers[0]['m.name']
final_answer = '{0}通常可以通过以下方式检查出来:{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
elif question_type == 'check_disease':
desc = [i['m.name'] for i in answers]
subject = answers[0]['n.name']
final_answer = '通常可以通过{0}检查出来的疾病有{1}'.format(subject, ';'.join(list(set(desc))[:self.num_limit]))
return final_answer
if __name__ == '__main__':
searcher = AnswerSearcher()
2.4.4 chatbot_graph.py 回答问题
#!/usr/bin/env python3
# coding: utf-8
from question_classifier import *
from question_parser import *
from answer_search import *
'''问答类'''
class ChatBotGraph:
def __init__(self):
self.classifier = QuestionClassifier()
self.parser = QuestionPaser()
self.searcher = AnswerSearcher()
def chat_main(self, sent):
answer = '您好,我是小勇医药智能助理,希望可以帮到您。如果没答上来,可联系https://liuhuanyong.github.io/。祝您身体棒棒!'
res_classify = self.classifier.classify(sent)
if not res_classify:
return answer
res_sql = self.parser.parser_main(res_classify)
final_answers = self.searcher.search_main(res_sql)
if not final_answers:
return answer
else:
return '\n'.join(final_answers)
if __name__ == '__main__':
handler = ChatBotGraph()
while 1:
question = input('用户:')
answer = handler.chat_main(question)
print('医生:', answer)
然后在chatbot_graph.py文件中进行语句整合,成功运行查询疾病健康。

参考文献
【1】https://blog.csdn.net/weixin_44023658/article/details/112503294?ops_request_misc=%257B%2522request%255Fid%2522%253A%2522166964389816782425686367%2522%252C%2522scm%2522%253A%252220140713.130102334…%2522%257D&request_id=166964389816782425686367&biz_id=0&utm_medium=distribute.pc_search_result.none-task-blog-2alltop_positive~default-1-112503294-null-null.142v67control,201v3add_ask,213v2t3_esquery_v2&utm_term=%E7%9F%A5%E8%AF%86%E5%9B%BE%E8%B0%B1&spm=1018.2226.3001.4187
【2】https://blog.csdn.net/zeroheitao/article/details/122925845
【3】