获取数据
library(jsonlite)
url <- 'https://view.inews.qq.com/g2/getOnsInfo?name=disease_h5'
pre <- fromJSON(url)
count <- fromJSON(pre$data)
#汇总
count$chinaTotal
$confirm
[1] 24433
$suspect
[1] 23260
$dead
[1] 493
$heal
[1] 968
#新增汇总
count$chinaAdd
$confirm
[1] 3962
$suspect
[1] 46
$dead
[1] 68
$heal
[1] 336
#每日汇总
count$chinaDayList
date confirm suspect dead heal
1 01.13 41 0 1 0
2 01.14 41 0 1 0
3 01.15 41 0 2 5
4 01.16 45 0 2 8
5 01.17 62 0 2 12
6 01.18 198 0 3 17
7 01.19 275 0 4 18
8 01.20 291 54 6 25
9 01.21 440 37 9 25
10 01.22 571 393 17 25
11 01.23 830 1072 25 34
12 01.24 1287 1965 41 38
13 01.25 1975 2684 56 49
14 01.26 2744 5794 80 51
15 01.27 4515 6973 106 60
16 01.28 5974 9239 132 103
17 01.29 7711 12167 170 124
18 01.30 9692 15238 213 171
19 01.31 11791 17988 259 243
20 02.01 14380 19544 304 328
21 02.02 17236 21558 361 475
22 02.03 20471 23214 425 632
23 02.04 24363 23260 491 892
#每日新增
count$chinaDayAddList
date confirm suspect dead heal
1 01.20 77 27 0 0
2 01.21 149 53 3 0
3 01.22 131 257 8 0
4 01.23 259 680 8 6
5 01.24 444 1118 16 3
6 01.25 688 1309 15 11
7 01.26 769 3806 24 2
8 01.27 1771 2077 26 9
9 01.28 1459 3248 26 43
10 01.29 1737 4148 38 21
11 01.30 1982 4812 43 47
12 01.31 2102 5019 46 72
13 02.01 2590 4562 45 85
14 02.02 2829 5173 57 147
15 02.03 3235 5072 64 157
16 02.04 3893 3971 65 262
数据可视化
新增趋势
add <- count$chinaDayAddList
#date转化为日期格式,其他转化为数字
add$date <- as.Date(add$date,'%m.%d')
add$confirm <- as.numeric(add$confirm)
add$suspect <- as.numeric(add$suspect)
#融合数据
library(reshape2)
melt_add <- melt(add,id='date',measure = c('confirm','suspect')
head(melt_add)
date variable value
1 2020-01-20 confirm 77
2 2020-01-21 confirm 149
3 2020-01-22 confirm 131
4 2020-01-23 confirm 259
5 2020-01-24 confirm 444
6 2020-01-25 confirm 688
#绘图
library(ggplot2)
library(ggthemes)
ggplot(melt_add) +
aes(x = date, y = value, colour = variable) +
geom_line(size = 1L) +
scale_color_hue() +
labs(x = " ", y = " ", title = "武汉肺炎疫情数据", subtitle = "新增趋势(人)", color = " ") +
theme_hc()+
scale_x_date(breaks = date,date_labels = '%m-%d')
累计确诊/疑似&累计治愈/死亡
daylist <- count$chinaDayList
daylist$date <- as.Date(daylist$date,'%m.%d')
daylist$confirm <- as.numeric(daylist$confirm)
daylist$suspect <- as.numeric(daylist$suspect)
daylist$dead <- as.numeric(daylist$dead)
daylist$heal <- as.numeric(daylist$heal)
#数据长宽转换
count_con_sus <- melt(daylist,id='date',measure = c('confirm','suspect'))
count_dead_heal <- melt(daylist,id='date',measure = c('dead','heal'))
#绘图
#累计确诊/疑似
ggplot(count_con_sus) +
aes(x = date, y = value, colour = variable) +
geom_line(size = 1L) +
scale_color_hue() +
labs(x = " ", y = " ", title = "武汉肺炎疫情数据", subtitle = "累计确诊/疑似(人)", color = " ") +
theme_hc()+
scale_x_date(date_labels = '%m-%d')
#累计治愈/死亡
ggplot(count_dead_heal) +
aes(x = date, y = value, colour = variable) +
geom_line(size = 1L) +
scale_color_hue() +
labs(x = " ", y = " ", title = "武汉肺炎疫情数据", subtitle = "累计治愈/死亡(人)", color = " ") +
theme_hc()+
scale_x_date(date_labels = '%m-%d')
GEO 可视化
GEO 可视化包有许多这里使用最简单的 REmap 包,它是基于百度 Echarts 的 R 语言包。
install.packages("devtools")
devtools::install_github("lchiffon/REmap")
library(REmap)
#json格式数据是嵌套的,可以用View函数依次查看全球、中国省级、中国市级情况
View(count$areaTree)
china <- count$areaTree[[2]][[1]]
china <- china[c(1,3)]
china$total <- china$total$confirm
china$total <- as.numeric(china$total)
remapC(china,color = 'orange',title = '武汉肺炎疫情地理可视化')
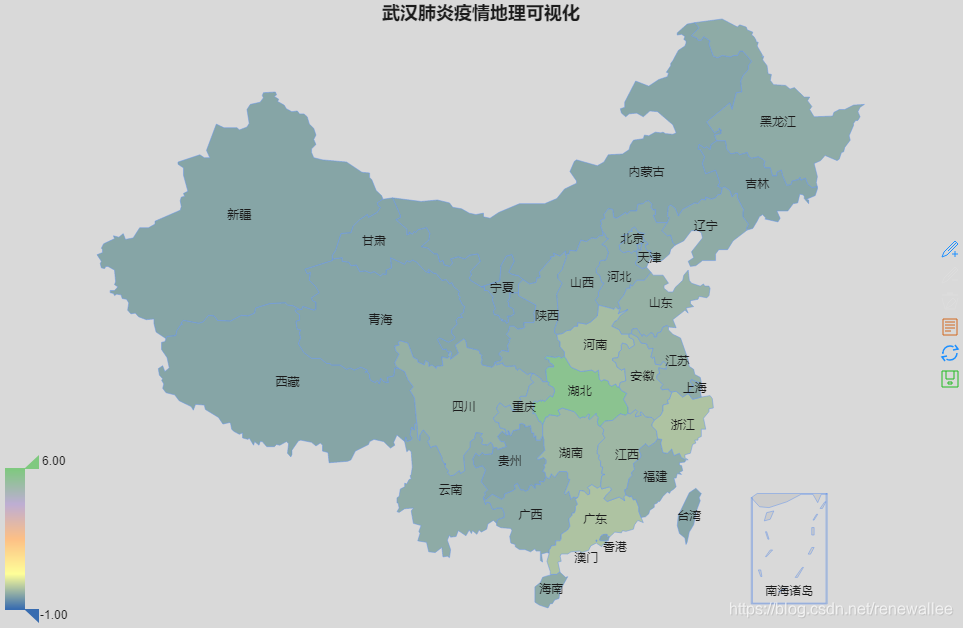
#省际差别大,地图区分不明显。尝试一下标准化数据
library(RColorBrewer)
col <- brewer.pal(5,'Accent')
china$total <- scale(china$total)
remapC(china,color = col,title = '武汉肺炎疫情地理可视化')