python机器学习读书笔记
- 读书笔记(六)Model Evaluation and Hyperparameter Tuning
读书笔记(六)Model Evaluation and Hyperparameter Tuning
嘿嘿嘿
一、Pipeline一体化操作
流程图如下

代码如下:
#The intermediate steps in a pipeline constitute scikit-learn transformers, and the last one is an estimator.
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
pipe_lr = make_pipeline(StandardScaler(),
PCA(n_components=2),
LogisticRegression(random_state=1,solver='lbfgs'))
pipe_lr.fit(X_train, y_train)#实际上调用了StandardScaler的fit_transform on the training data,
#并且将标准化后的参数传递给PCA,PCA操作同理
y_pred = pipe_lr.predict(X_test)#LogisticRegression的操作
print('Test Accuracy: %.3f' % pipe_lr.score(X_test, y_test))
二、Holdout cross-validation and K-fold cross-validation
1. Holdout cross-validation
基本方法:
将data分为三个部分,分别为 training set、validation set 、test set
训练集用于训练不同的模型,验证集用于模型选择,而测试集则用于 obtain a less biased estimate of its ablility to generalize data. 测试模型的泛化能力。
缺点:
结果对于dataset的三个subset的划分十分敏感,not robust enough~~(相对于下面将要介绍的K-fold cross-validation )
如图:

2.K-fold cross-validation
基本方法:
将dataset不放回地随机分为k组,其中k-1组用于training,剩下1组用于testing。重复该步骤k次,得到k个model estimates。计算出平均值,可以得到一个比Holdout cross-validation更不依赖于data的分割的估计。
如图:

优点:
每一个样本点都有且仅有一次成为training set 和 test set的一部分。相较于holdout方法,减小了过拟合的可能(lower-variance estimate)
小注意~:
-
1.这个一般方法用于调参,调参完毕,再将整个dataset代入模型之中得到一个最后的performance estimate.
-
2.k的一般默认取值为10,对于较小的数据集,我们可以适当增大k,由于更多的数据能被用上,这样可以lower bias(但是增大了计算量并且可能导致higher variance)。对于较大的数据集,我们可以适当地减小k(e.g. k
=5),相对地可以减少计算量。 -
#在数据集极端小的情况下,一个Kfold的极端例子是k=n,称为leave-one-out(LOO),即每次test sample只有一个样本点。
3.Stratified k-fold(分层)
基本方法:
In stratified cross-validation, the calss proportions are preserved in each fold to ensure that each fold is representative of the class proportions in the training dataset.(即每一个fold中不同类的比例与总体保持一致,故称为stratified,不禁让人联想到train_test_split()中的参数,分层取样)
import numpy as np
from sklearn.model_selection import StratifiedKFold
kfold = StratifiedKFold(n_splits=10,
random_state=1).split(X_train, y_train)
scores = []
for k, (train, test) in enumerate(kfold):
pipe_lr.fit(X_train[train], y_train[train])
score = pipe_lr.score(X_train[test], y_train[test])
scores.append(score)
print('Fold: %2d, Class dist.: %s, Acc: %.3f' % (k+1,
np.bincount(y_train[train]), score))
print('\nCV accuracy: %.3f +/- %.3f' % (np.mean(scores), np.std(scores)))
结果如下
Fold: 1, Class dist.: [256 153], Acc: 0.935
Fold: 2, Class dist.: [256 153], Acc: 0.935
Fold: 3, Class dist.: [256 153], Acc: 0.957
Fold: 4, Class dist.: [256 153], Acc: 0.957
Fold: 5, Class dist.: [256 153], Acc: 0.935
Fold: 6, Class dist.: [257 153], Acc: 0.956
Fold: 7, Class dist.: [257 153], Acc: 0.978
Fold: 8, Class dist.: [257 153], Acc: 0.933
Fold: 9, Class dist.: [257 153], Acc: 0.956
Fold: 10, Class dist.: [257 153], Acc: 0.956
CV accuracy: 0.950 +/- 0.014
一个更为简洁的方式using stratified k-fold cross-validation
#(其实我个人感觉参数input里也妹有分层的参数鸭,但是查了github上源代码人家好像default就都是stratified的,好吧。。。。ヽ(ー_ー)ノ)
from sklearn.model_selection import cross_val_score
scores = cross_val_score(estimator=pipe_lr,
X=X_train,
y=y_train,
cv=10,
n_jobs=-1)#n_jobs代表CPU的核心数,其中-1代表使用所有CPU
print('CV accuracy scores: %s' % scores)
print('CV accuracy: %.3f +/- %.3f' % (np.mean(scores), np.std(scores)))
结果如下:
CV accuracy scores: [0.93478261 0.93478261 0.95652174 0.95652174 0.93478261 0.95555556
0.97777778 0.93333333 0.95555556 0.95555556]
CV accuracy: 0.950 +/- 0.014
三、Diagnose bias and variance problems with learning curves
1. illustration of bias and variance
首先来看一幅图:
总体来说,
-
当一个模型相对于训练集来说太过于复杂—(there are too many degrees of freedom or parameters in the model),模型就会趋于过拟合而缺乏泛化能力,因而在测试集或验证集上表现不佳。从图像上来看,即蓝线代表的Validation accuracy 始终在 Training Accuracy 之下。
-
通常来说,增加Number of training sample 常常可以减小过拟合的程度。从图像上来看,不管是哪种图像,当横坐标增加,两条曲线趋与一致。但是实践中这常常不现实或者不经济。

左上:Underfit(high bias)
模型的Training and Validation accuracy 都较低,说明模型欠拟合。此时需要增加模型的参数以增加模型的复杂度。例如:收集或者构造更多的特征,或者减小正则化的程度(C or λ)。
右上:Overfit (high variance)
模型的Training and Validation accuracy 之间有着一个较大的gap,说明过拟合。(training的预测能力很好,但是相对地validation的预测能力很差,中间有着巨大的差距,且此差距随着样本数的增加而减小)。此时需要减小模型的复杂度或者是收集更多的数据集。例如:增加正则化的程度,SBS特征选取,PCA 或者 LDA 特征提取。
用自己的数据集画一个实际的图:
import matplotlib.pyplot as plt
from sklearn.model_selection import learning_curve
pipe_lr = make_pipeline(StandardScaler(),
LogisticRegression(penalty='l2', random_state=1,solver = 'lbfgs'))
#train_sizes返回的是实际的分组数量,注意与参数中的区分开
train_sizes, train_scores, test_scores =\
learning_curve(estimator=pipe_lr,
X=X_train,
y=y_train,
train_sizes=np.linspace(0.1, 1.0, 10),
cv=10,
n_jobs=1)
train_mean = np.mean(train_scores, axis=1)
train_std = np.std(train_scores, axis=1)
#计算出标准差好画带状图
test_mean = np.mean(test_scores, axis=1)
test_std = np.std(test_scores, axis=1)
plt.plot(train_sizes, train_mean,
color='blue', marker='o',
markersize=5, label='training accuracy')
#画带状图
plt.fill_between(train_sizes,
train_mean + train_std,
train_mean - train_std,
alpha=0.15, color='blue')
plt.plot(train_sizes, test_mean,
color='green', linestyle='--',
marker='s', markersize=5,
label='validation accuracy')
plt.fill_between(train_sizes,
test_mean + test_std,
test_mean - test_std,
alpha=0.15, color='green')
plt.grid()
plt.xlabel('Number of training samples')
plt.ylabel('Accuracy')
plt.legend(loc='lower right')
plt.ylim([0.8, 1.03])
plt.tight_layout()
#plt.savefig('images/06_05.png', dpi=300)
plt.show()
如图

四、Addressing overfitting and underfitting with validation curves
1. 初级版:用sklearn中的validation_curve调参
不同于上述以样本量为自变量的图,在此处,我们将重点放在如何调整参数使得从而提高validation accuracy。因此我们可以对上述代码稍作改动,在确定的样本数量下,将inverse 正则化参数C 作为自变量画图如下:
from sklearn.model_selection import validation_curve
param_range = [0.001, 0.01, 0.1, 1.0, 10.0, 100.0]
train_scores, test_scores = validation_curve(
estimator=pipe_lr,
X=X_train,
y=y_train,
param_name='logisticregression__C',
param_range=param_range,
cv=10)
train_mean = np.mean(train_scores, axis=1)
train_std = np.std(train_scores, axis=1)
test_mean = np.mean(test_scores, axis=1)
test_std = np.std(test_scores, axis=1)
plt.plot(param_range, train_mean,
color='blue', marker='o',
markersize=5, label='training accuracy')
plt.fill_between(param_range, train_mean + train_std,
train_mean - train_std, alpha=0.15,
color='blue')
plt.plot(param_range, test_mean,
color='green', linestyle='--',
marker='s', markersize=5,
label='validation accuracy')
plt.fill_between(param_range,
test_mean + test_std,
test_mean - test_std,
alpha=0.15, color='green')
plt.grid()
plt.xscale('log')
plt.legend(loc='lower right')
plt.xlabel('Parameter C')
plt.ylabel('Accuracy')
plt.ylim([0.8, 1.0])
plt.tight_layout()
# plt.savefig('images/06_06.png', dpi=300)
plt.show()
如图所示

可以看出,当C较小时,即正则化力度较大,模型欠拟合。当C不断增大的时候,即意味着正则化力度减小,模型有过拟合的迹象,此时sweet spot 为C = 1.
2. 升级版:用grid search 调参
首先认识一下机器学习中的两种参数:第一中是通过training data 确定的参数,比如逻辑回归中的权重;第二种是我们自己设置的,不受训练过程影响的参数,即超参数(Hyperparameters or tuning parameter)比如逻辑回归中的正则化参数与决策树中树的深度。
grid search 其实没有什么高深之处,it’s a brute-force exhaustive search paradiam. 就是尝试所有的参数组合,找出最优的那一组。
代码如下:
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
pipe_svc = make_pipeline(StandardScaler(),
SVC(random_state=1))
param_range = [0.0001, 0.01, 0.1, 1.0, 10.0, 100.0, 1000.0]
param_grid = [{'svc__C': param_range,
'svc__kernel': ['linear']},
{'svc__C': param_range,
'svc__gamma': param_range,
'svc__kernel': ['rbf']}]
gs = GridSearchCV(estimator=pipe_svc,
param_grid=param_grid,
scoring='accuracy',
cv=10,
n_jobs=-1)
gs = gs.fit(X_train, y_train)
print(gs.best_score_)
print(gs.best_params_)
结果如下:
0.9846153846153847
{'svc__C': 1000.0, 'svc__gamma': 0.0001, 'svc__kernel': 'rbf'}
最后我们再将测试集代入最优模型之中
代码如下:
clf = gs.best_estimator_
# clf.fit(X_train, y_train)
# Note that the line above is not necessary, because
# the best_estimator_ will already be refit to the complete
# training set because of the refit=True setting in GridSearchCV
# (refit=True by default). Thanks to a reader, German Martinez,
# for pointing it out.
#这个德国哥们儿有点nb,甚至能改错
print('Test accuracy: %.3f' % clf.score(X_test, y_test))
结果如下:
Test accuracy: 0.974
3. Nested cross-validation
选择算法用,在一项实验中,Varma and Simon conclude that the true error of the estimate is almost unbiased relative to the test set when nested cross-validation is used.(具体原因也不知道是为啥,嗯,反正用于比较算法就对了)
具体操作:

内层交叉验证(innner loop):用于模型选择,可以进行特征工程处理数据。
外层交叉验证(outer loop):用于模型评估,使用所有数据集进行分割,而不仅是训练集,且用Stratified K-Fold保证类别比例不变。外层每一折都使用内层得到的最优参数组合进行训练。
#此处参考↑:
#作者:行走的程序猿
#链接:https://www.jianshu.com/p/cdf6df99b44b
下面我们来比较SVM与决策树模型的表现:
sklearn实现代码如下
1.SVM
gs = GridSearchCV(estimator=pipe_svc,
param_grid=param_grid,
scoring='accuracy',
cv=2)
scores = cross_val_score(gs, X_train, y_train,
scoring='accuracy', cv=5)
print('CV accuracy: %.3f +/- %.3f' % (np.mean(scores),
np.std(scores)))
结果如下:
CV accuracy: 0.974 +/- 0.015
2.决策树
from sklearn.tree import DecisionTreeClassifier
gs = GridSearchCV(estimator=DecisionTreeClassifier(random_state=0),
param_grid=[{'max_depth': [1, 2, 3, 4, 5, 6, 7, None]}],
scoring='accuracy',
cv=2)
scores = cross_val_score(gs, X_train, y_train,
scoring='accuracy', cv=5)
print('CV accuracy: %.3f +/- %.3f' % (np.mean(scores),
np.std(scores)))
结果如下:
CV accuracy: 0.934 +/- 0.016
五、Performance evaluation matrix
图像解释很清楚~~

sklearn 中已经封装好的挺方便
from sklearn.metrics import confusion_matrix
pipe_svc.fit(X_train, y_train)
y_pred = pipe_svc.predict(X_test)
confmat = confusion_matrix(y_true=y_test, y_pred=y_pred)
print(confmat)
如下:
[[71 1]
[ 2 40]]
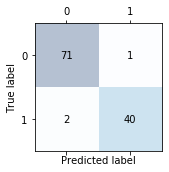
美化一下:
fig, ax = plt.subplots(figsize=(2.5, 2.5))
ax.matshow(confmat, cmap=plt.cm.Blues, alpha=0.3)
for i in range(confmat.shape[0]):
for j in range(confmat.shape[1]):
ax.text(x=j, y=i, s=confmat[i, j], va='center', ha='center')
plt.xlabel('Predicted label')
plt.ylabel('True label')
plt.tight_layout()
#plt.savefig('images/06_09.png', dpi=300)
plt.show()
from sklearn.metrics import precision_score, recall_score, f1_score
print('Precision: %.3f' % precision_score(y_true=y_test, y_pred=y_pred))
print('Recall: %.3f' % recall_score(y_true=y_test, y_pred=y_pred))
print('F1: %.3f' % f1_score(y_true=y_test, y_pred=y_pred))
Precision: 0.976
Recall: 0.952
F1: 0.964
可以更换上述grid_search的评价指标了
#以f1_score为指标,以0为positive
scorer = make_scorer(f1_score, pos_label=0)
from sklearn.metrics import make_scorer
scorer = make_scorer(f1_score, pos_label=0)
c_gamma_range = [0.01, 0.1, 1.0, 10.0]
param_grid = [{'svc__C': c_gamma_range,
'svc__kernel': ['linear']},
{'svc__C': c_gamma_range,
'svc__gamma': c_gamma_range,
'svc__kernel': ['rbf']}]
gs = GridSearchCV(estimator=pipe_svc,
param_grid=param_grid,
scoring=scorer,
cv=10,
n_jobs=-1)
gs = gs.fit(X_train, y_train)
print(gs.best_score_)
print(gs.best_params_)
0.9862021456964396
{'svc__C': 10.0, 'svc__gamma': 0.01, 'svc__kernel': 'rbf'}
六、ROC曲线(Receiver Operating Characteristics)
以FPR与TPR分别为横纵坐标绘制ROC曲线:
from sklearn.metrics import roc_curve, auc
from scipy import interp
pipe_lr = make_pipeline(StandardScaler(),
PCA(n_components=2),
LogisticRegression(penalty='l2',
random_state=1,
C=100.0))
X_train2 = X_train[:, [4, 14]]
cv = list(StratifiedKFold(n_splits=3,
random_state=1).split(X_train, y_train))
fig = plt.figure(figsize=(7, 5))
mean_tpr = 0.0
mean_fpr = np.linspace(0, 1, 100)
all_tpr = []
for i, (train, test) in enumerate(cv):
probas = pipe_lr.fit(X_train2[train],
y_train[train]).predict_proba(X_train2[test])
fpr, tpr, thresholds = roc_curve(y_train[test],
probas[:, 1],
pos_label=1)
mean_tpr += interp(mean_fpr, fpr, tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr, tpr)
plt.plot(fpr,
tpr,
label='ROC fold %d (area = %0.2f)'
% (i+1, roc_auc))
plt.plot([0, 1],
[0, 1],
linestyle='--',
color=(0.6, 0.6, 0.6),
label='random guessing')
mean_tpr /= len(cv)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
plt.plot(mean_fpr, mean_tpr, 'k--',
label='mean ROC (area = %0.2f)' % mean_auc, lw=2)
plt.plot([0, 0, 1],
[0, 1, 1],
linestyle=':',
color='black',
label='perfect performance')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('false positive rate')
plt.ylabel('true positive rate')
plt.legend(loc="lower right")
plt.tight_layout()
# plt.savefig('images/06_10.png', dpi=300)
plt.show()

